Arbabshirani MR, Plis S, Sui J, Calhoun VD.
Blog post written by: Andres Rodrigo Zapata Rodriguez
Introduction
Single Subject prediction of Brain disorders has always been an attractive field for researchers, especially considering the emerging fields of Neuroimaging (NI) and machine learning. NI is an incentive for many scientists of several heterogeneous domains to enter the study of the human brain. Different imaging modalities (sMRI, fMRI, dMRI) bring along different non-invasive perspectives on the brain. Unfortunately, many diseases have no clinical tests for their identification. The promise of the combination of Neurosciensce(NS) NI and Machine Learning(ML)l is to cast light on these less-understood diseases and be able to treat them in an informed manner. Although, in recent times, this topic has been crowded with deep learning approaches, this has not always been the case.
The presented survey was conducted precisely at this time - at the advent of deep learning. It treats a wide array of methodologies within ML from those times that supersede today’s mostly deep learning-focused era. The survey has crystallized three crucial points. First, it supplies a good overview of the research area and delineates its trends. Secondly, it identifies the common pitfalls in this area. Thirdly it determines the shortcomings and emerging trends in that era.
Background
Before getting into the central part of the survey [1], two important background topics can be helpful in understanding the ideas put forth in this survey.
Imaging techniques and Diseases
Firstly, having a notion of the nature of the imaging techniques and the diseases they shed light on is helpful for grasping the underlying problem of this research domain. In the two graphs below, we have a short overview of these topics with suggestions for reading material.
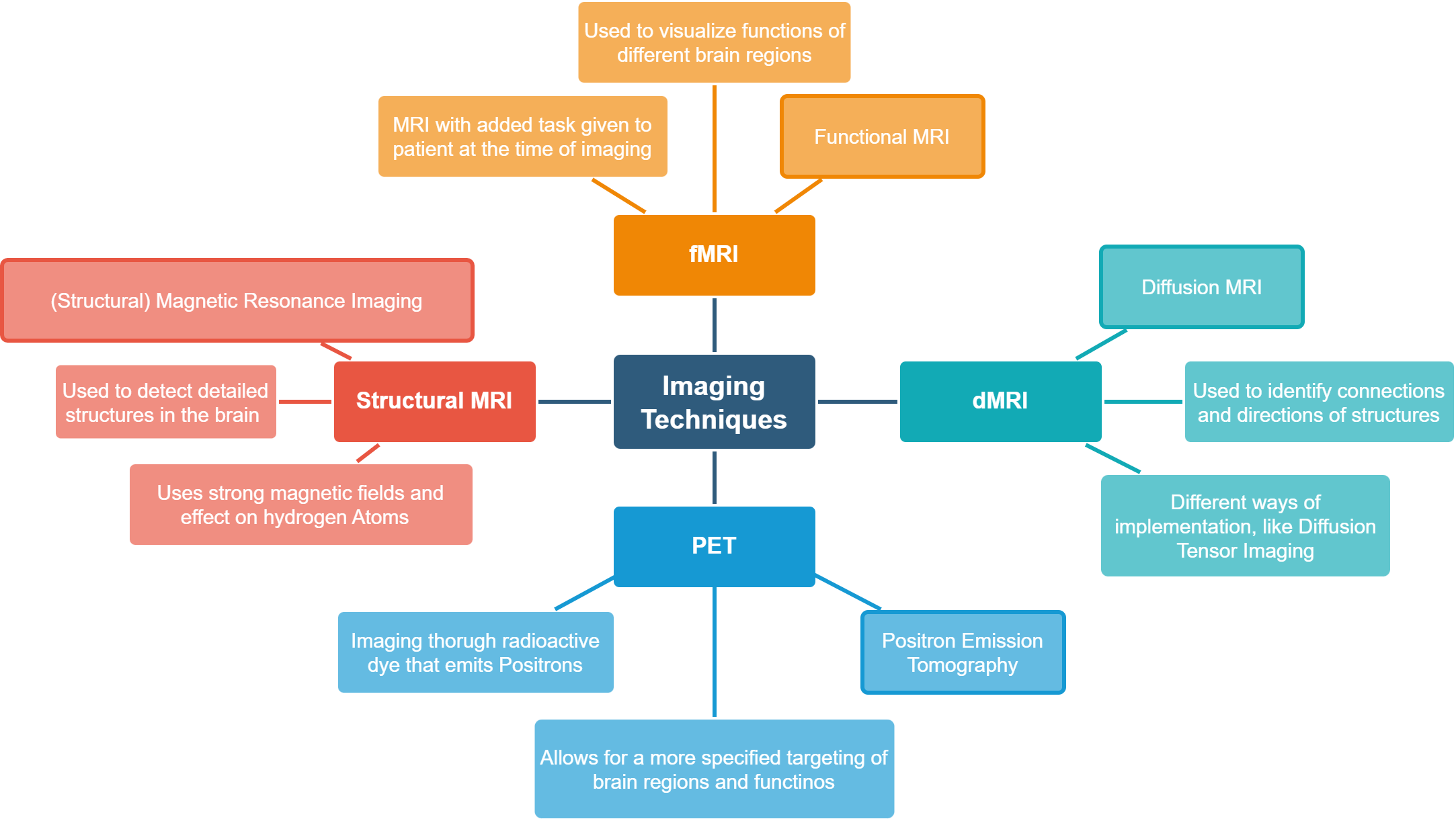
Fig. 1 Imaging Techniques [6] [7]
Fig. 2 Diseases [5]
Group difference vs. Classification
Secondly, this survey points out a significant difference in how we analyze the raw data against how we use the outcoming analysis of our data that brings along a challenge that we may not assume: Findings in the studies until the time of the survey were helpful at finding and studying biomarkers but are not suited for the clinical diagnostic or prognostic scenario, where individual discrimination power of the biomarkers would be required. One of the reasons behind this is the difficulty of translating information from a Group difference to a classification task. There are two ways of viewing medical data. In the first perspective, one tries to identify the relationship of specific values within or among groups. In the second one, the goal is to classify one subject into one of the different categories[2]. This study conveys the idea that results from a group-level analysis are not translatable or even usable in the domain of single-subject classification
Overfitting
Overfitting is a substantial issue when dealing with NI datasets because of the very scarce available data. Factoring into this issue is that most NI datasets are composed of big and complex samples. As a solution, the survey points to the standard deep learning strategies against overfitting and states that a good feature selection is also an excellent measure to prevent overfitting. The prevalence of this problem in specifically this research environment can be seen as a comorbidity of the fact that there is a limited sample size.[3]
Reporting classification results
The survey advocates for presenting a detailed result of the classification in the form of the confusion table and related Performance measures. In addition, the survey points explicitly out two common reporting issues encountered in works related to single-subject prediction with NI data. The first problem is that sometimes reporting the overall accuracy is not representative of the result because the class distributions need to be balanced. Here the survey suggests using the F1 score or balanced versions of the standard performance measures. The second common reporting issue is using the random chance to set the bar for performance.
Comparison of accuracies across studies
In literature, it is common to claim to outperform other studies only by using overall accuracy; however, it is a very reductive mode of comparison since ML applications are influenced by a vast array of variables - starting from the obvious hyperparameters and finishing with more hidden
aspects such as the abstract design of the study itself. Therefore a more in-depth comparison should be taken into account, starting from the considerations of the previous section.
Hyperparameter optimization
Hyperparameter optimization is the search for optimal values for a ML architecture for a specific task at hand. This search impacts the final performance of the considered implementation significantly. Differences in Hyperparameter optimization can lead to substantial differences in performance, even within the most straightforward architectures of the same type. This makes comparing between studies even more difficult since they can have different levels, forms, and even approaches to Hyperparameter optimization, which are also not reported in many cases.
Operating on decentralized data
The authors of the survey argue that a suitable solution for the small globally available sample sizes is to use algorithms implemented to work on data distributed across research groups in a decentralized manner. These algorithms are planned to have privacy-conserving measures implemented in them and would allow for a more efficient and more suitable framework for dealing with sensitive data in the age of big data. The general goal of these approaches is to preserve the information and efficacy of the given data and minimize the amount of data that has to be shared. Pragmatically speaking, the privacy-providing method suggested and deemed as suitable for the data at hand is the e-differential privacy model since it is a model concerned with sharing a function of the data and not the data itself. Furthermore, privacy is increasingly important because of the envisioned decentralized approach preached by the survey authors. The combination of these two methods, decentralized algorithms, and e-differential privacy, is favorable because it mitigates the loss of quality in data due to noise instruction through e-differentially privacy.[4]
Multimodal neuroimaging studies
Utilizing multiple modalities has been shown to increase confidence in the conclusions deducted from the algorithms that implement them. The survey points out that most of these studies favor one modality over the other and there is no fully integrated fusion of the data. Single-subject prediction using multimodal NI data In the times of the survey, it was shown that having a multimodal view of brain disorders reveals distinctive patterns that are not visible when only considering a unimodal view of the patient. This makes the use of a unimodal view in single-subject prediction a logical step in research, which could finally lead to a faster and more precise diagnostication process for the patients. [8] [9]
Deep learning in Neuroimaging
As of the survey’s authors, two main characteristics of deep learning have made it attractive for researchers. The first enticing feature is that deep learning is a data-driven technique that learns useful features automatically. This works against the issue of having to manually select features in a vast space using insufficient univariate statistical tests. The second characteristic is the non-linear nature of deep learning methods, which can also approximate highly non-linear data. This allows deep learning methods to learn models with significantly fewer parameters.
Standard machine learning competitions in Neuroimaging
ML research has profited hugely from standard competitions where participants are provided a labeled training dataset and an unlabeled test dataset. The participants profit from a standard dataset that is already prepossessed to some degree, thus allowing researchers from other non-neuroscience domains to enter the competition with less effort. This competition also brings along the exploration of a more brought array of architectures because it attracts researchers from many different fields, thus expanding the solution approaches explored in the field. Unfortunately, the small globally available. dataset combined with the privacy concerns linked to the medical data of humans make these kinds of competitions more difficult and scarce for NI data.
Discussion
Previous single-subject prediction surveys
In the past several surveys on single-subject prediction have been published. However, these were usually limited to one disease’s scope or specific technique. Also, the past surveys are outdated. Therefore the considered survey has been identified by the authors to be the most comprehensive one at the time of its release.
Meaning Today
After having gone through the outcomes of this survey, a question arises: what meaning do the outcomes of this survey have at the current time? One might say that there is no interest in older approaches; however, the knowledge about more traditional and maybe more understandable approaches can bring substantial prof- its to research at the cutting edge of science. Notably, it is valuable to learn from the learned lessons of researchers in the past, and this(but not exclusively) makes this paper valuable for readers today. Additionally, past architectures should be taken as a baseline to which modern techniques have to compare against. Also, this survey has shown us that a substantial amount of effort has been applied to Alzheimer’s disease research, while other areas have suffered a lack of focus. This vast amount of experience and knowledge could now be exploited by being translated and ported to other fields of interest. Considering the identified five Pitfalls in ML in NI, we can say that most of them are still a problem in the research community; however, since most of the publications utilize deep learning techniques, it is possible to say that the issue of feature selection bias is less of a problem today. From the shortcomings and Emerging trends identified by the survey, all are more or less true up to the current times. Both shortcomings: lack of data, and the difficulty of decease sub-type classification and differential diagnosis, are still roadblocks in research in this area. What the emergent trends concern, only deep learning has truly been explored thoroughly in the last years, while the other emergent trends still promise to bring fruits in the future.
Reference
[1] Arbabshirani MR, Plis S, Sui J, Calhoun VD. Single subject prediction of brain disorders in neuroimaging: Promises and pitfalls. Neuroimage. 2017 Jan 15;145(Pt B):137-165. doi: 10.1016/j.neuroimage.2016.02.079. Epub 2016 Mar 21. PMID: 27012503; PMCID: PMC5031516.
[2]Bishop, C.~.M., 2006. Pattern Recognition and Machine Learning (Information Science and 1107 Statistics). Springer.
[3] Plis, S.M., Hjelm, D., Salakhutdinov, R., Allen, E.A., Bockholt, H.J., Long, J.D., Johnson, H.J., Paulsen, J., Turner, J.A., Calhoun, V.D., 2014. Deep learning for neuroimaging: a validation study. Front. Neurosci. 8.
[4] Sarwate, A.D., Plis, S.M., Turner, J.A., Arbabshirani, M.R., Calhoun, V.D., 2014. Sharing privacy-sensitive access to neuroimaging and genetics data: a review and preliminary validation. Front. Neuroinform. 8, 35. http://dx.doi.org/10.3389/fninf.2014.00035.
[5] FRIEDMAN, Howard S. Encyclopedia of mental health. Academic Press, 2015.
[6] A. Lenartowicz, R.A. Poldrack, Brain Imaging, Editor(s): Sergio Della Sala, Encyclopedia of Behavioral Neuroscience, 2nd edition (Second Edition), Elsevier, 2017,Pages 77-83,ISBN 9780128216361, https://doi.org/10.1016/B978-0-12-809324-5.00274-1.
[7] Andreas Otte, Ulrike Halsband, Brain imaging tools in neurosciences, Journal of Physiology-Paris, Volume 99, Issues 4–6, 2006, Pages 281-292, ISSN 0928-4257, https://doi.org/10.1016/j.jphysparis.2006.03.011.
[8] Rashmin Achalia, Anannya Sinha, Arpitha Jacob, Garimaa Achalia, Varsha Kaginalkar, Ganesan Venkatasubramanian, Naren P. Rao, A proof of concept machine learning analysis using multimodal neuroimaging and neurocognitive measures as predictive biomarker in bipolar disorder, Asian Journal of Psychiatry, Volume 50, 2020, 101984, ISSN 1876-2018, https://doi.org/10.1016/j.ajp.2020.101984.
[9] Tulay, Emine Elif, et al. "Multimodal neuroimaging: basic concepts and classification of neuropsychiatric diseases." Clinical EEG and neuroscience 50.1 (2019): 20-33.


